叶绿体基因组测序分析
-
叶绿体基因组测序分析
介绍:叶绿体中包含的所有遗传物质称为叶绿体基因组。叶绿体基因组包含大量的遗传信息,同时叶绿体基因组结构保守,除了反向重复区外所有基因均为单拷贝,几乎不存在(或很少受)旁系同源基因的干扰,同时几乎没有来自线粒体和核的外源DNA。此外,叶绿体DNA的核苷酸替代率适中,在应用上有很大的价值,而且叶绿体基因组编码区和非编码区分子进化速率差异较大(前者慢后者快),因此可分别为不同层级的系统学研究提供依据。
随着测序技术的发展,下一代测序技术(NGS)使基因组的测序不入了一个低成本及高通量测序的时代。基于叶绿体基因组的高度保守性及二代测序的高通量性,研究者可以不通过分离纯化叶绿体DNA,而直接从全基因组高通量测序数据中获取叶绿体基因组。根据我司数百例叶绿体基因组组装经验,只有极少数类群需要结合三代测序或一代测序补gap。
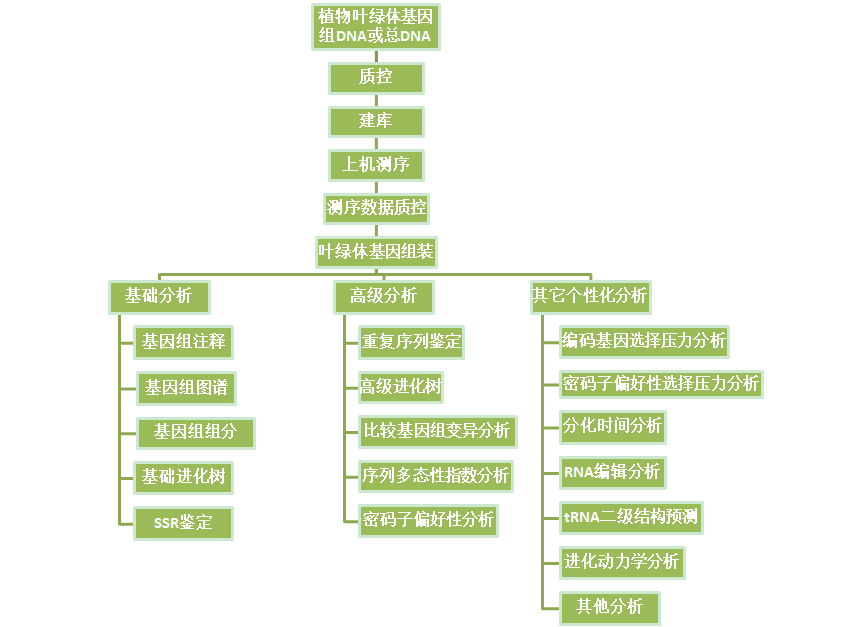
叶绿体基因组测序分析流程图
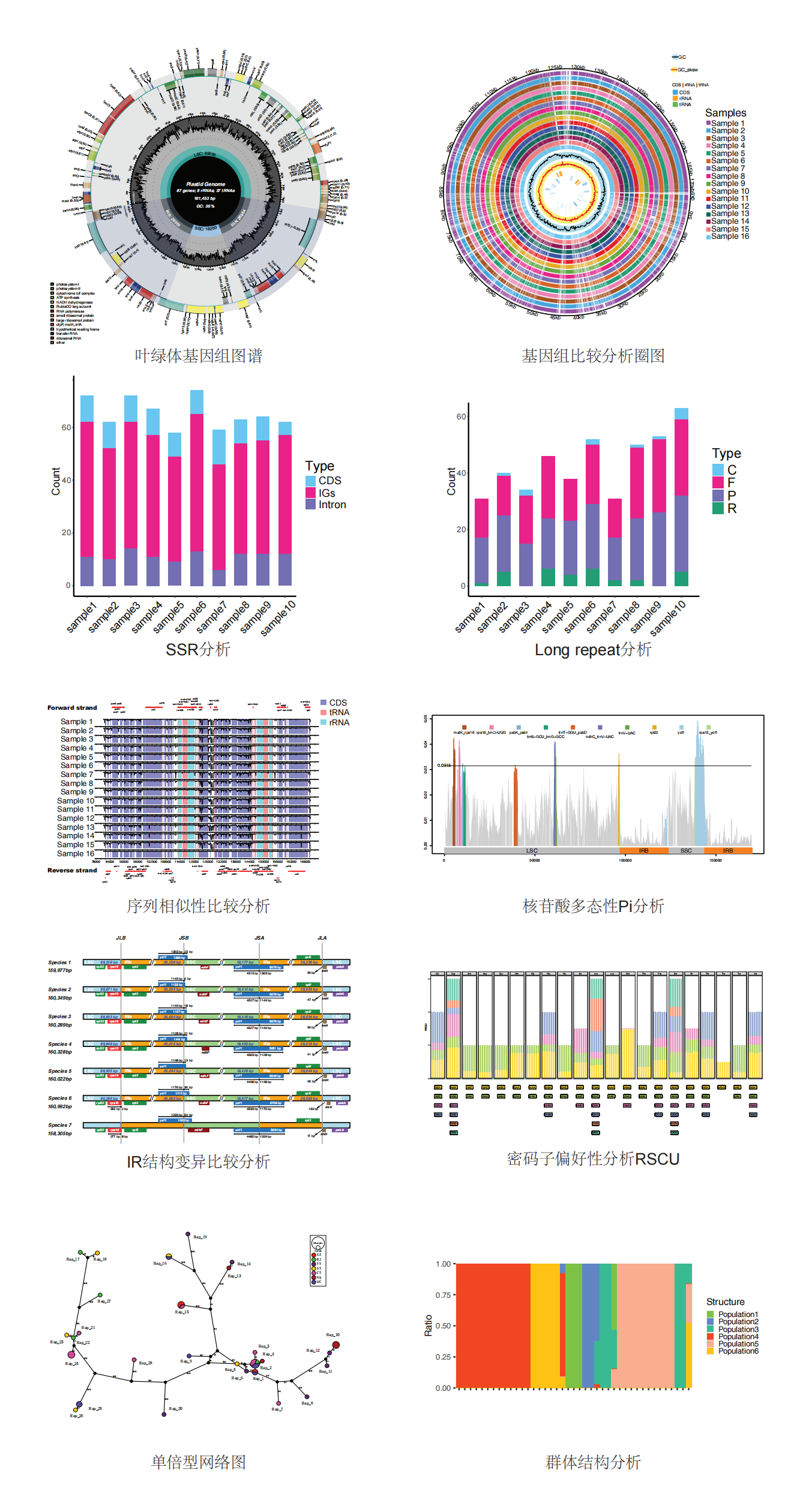
叶绿体分析主要内容
代表文章(#为我司技术人员为共同作者):
李国良, 张鸿, 林赵淼, 等. 叶菜型甘薯叶绿体基因组及其特征分析[J]. 西南大学学报 (自然科学版), 2023, 45(10): 43-53.
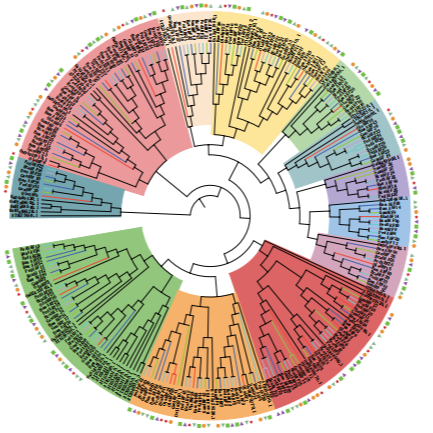
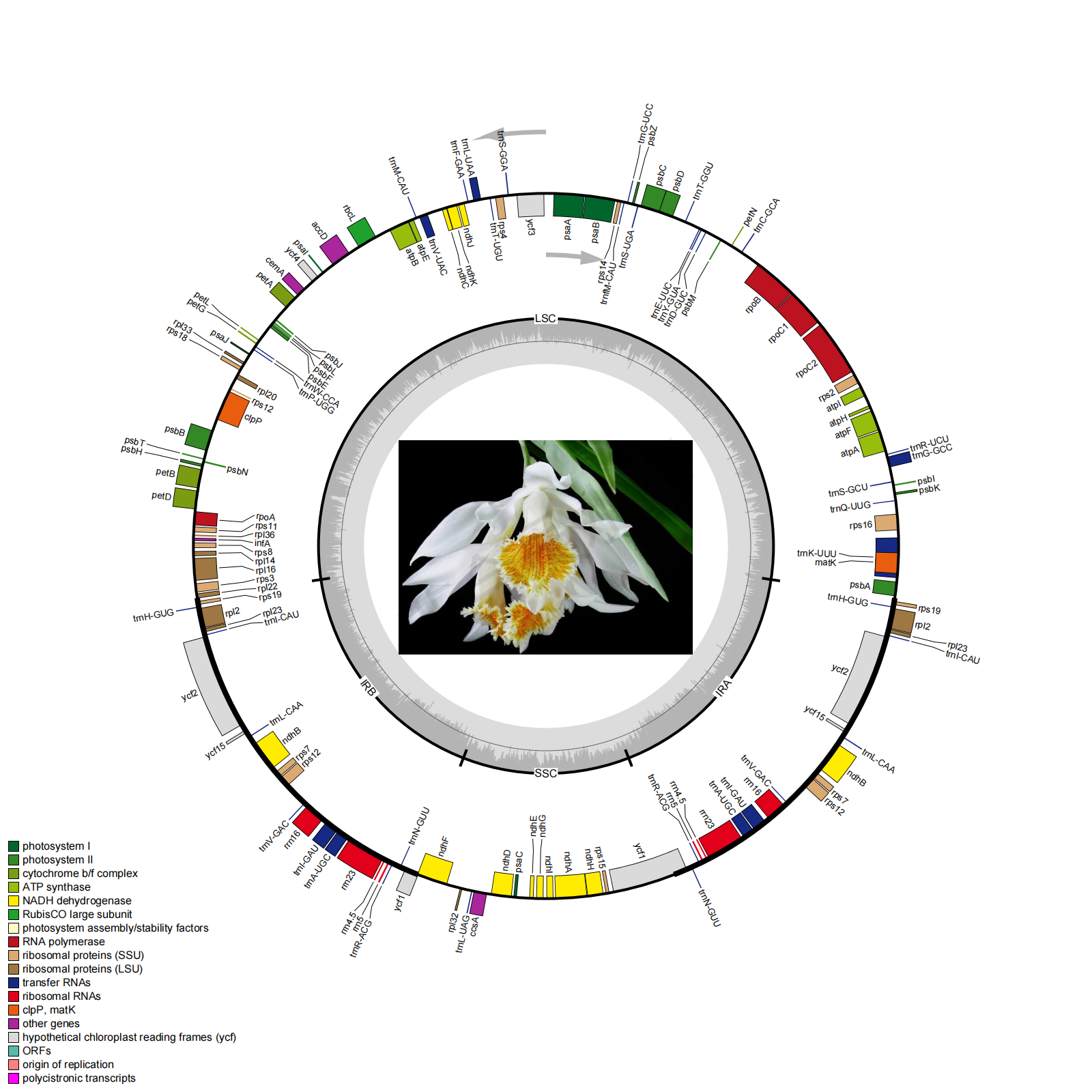
Li G L, Zhang H, Lin Z, et al. Comparative analysis of chloroplast and mitochondrial genomes of sweet potato provides evidence of gene transfer[J]. Scientific Reports, 2024, 14(1): 4547.Li, L., Wu, Q., Zhai, J. et al. Comparative chloroplast genomics of 24 species shed light on the genome evolution and phylogeny of subtribe Coelogyninae (Orchidaceae). BMC Plant Biol 24, 31 (2024). https://doi.org/10.1186/s12870-023-04665-2。
Yue M, Chen H, Lei X, et al. Novel molecular markers for Taxodium breeding from the chloroplast genomes of four artificial Taxodium hybrids[J]. Frontiers in Genetics, 14: 1193023.
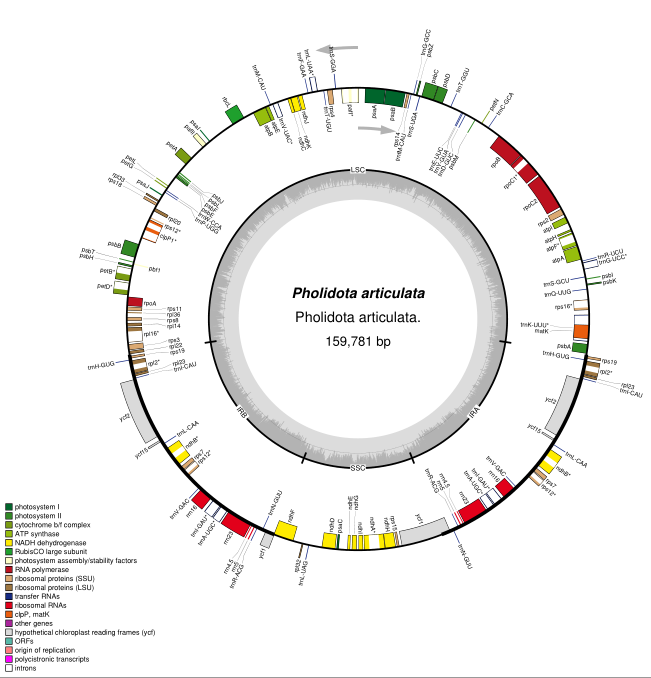
Li L, Wang W, Zhang G, et al. Comparative analyses and phylogenetic relationships of thirteen Pholidota species (Orchidaceae) inferred from complete chloroplast genomes. BMC Plant Biol 23, 269 (2023). https://doi.org/10.1186/s12870-023-04233-8
Ye Y, Liu J, Zhou Y, Zhu G, Tan J, Xu Y. Complete Chloroplast Genome Sequences of Four Species in the Caladium Genus: Comparative and Phylogenetic Analyses. Genes 2022, 13, 2180. https://doi.org/10.3390/genes13122180
#Chong, X., et al., Comparative chloroplast genome analysis of 10 Ilex species and the development of species-specific identification markers. Industrial Crops and Products, 2022. 187: p. 115408.
#Li L, Wu Q, Fang L, et al. Comparative Chloroplast Genomics and Phylogenetic Analysis of Thuniopsis and Closely Related Genera within Coelogyninae (Orchidaceae)[J]. Frontiers in Genetics, 2022: 505.
#Yi, S.; Lu, H.; Wang, W.; Wang, G.; Xu, T.; Li, M.; Gu, F.; Chen, C.; Han, B.; Liu, D. The Chloroplast Genome of Wild Saposhnikovia divaricata: Genomic Features, Comparative Analysis, and Phylogenetic Relationships. Genes 2022, 13, 931.
#Liu Q, Li X, Li M, et al. Comparative chloroplast genome analyses of Avena: insights into evolutionary dynamics and phylogeny[J]. BMC Plant Biology, 2020, 20(1): 1-20.
#Duan, H., Guo, J., Xuan, L. et al. Comparative chloroplast genomics of the genus Taxodium. BMC Genomics 21, 114 (2020).
#Li X, Li Y, Zang M, et al. Complete Chloroplast Genome Sequence and Phylogenetic Analysis of Quercus acutissima[J]. International Journal of Molecular Sciences. 2018, 19(8): 2443. IF=3.687
Wang L, Yu X, Xu W, et al. Complete chloroplast genome sequencing support Angelica decursiva is an independent species from Peucedanum praeruptorum[J]. Physiology and Molecular Biology of Plants, 2021: 1-13.
Liu X, Sun C, Li M, et al. The complete chloroplast genome sequence of Actinidia chinensis Planch.‘Hongyang’, a typical red core pulp in China[J]. Mitochondrial DNA Part B, 2022, 7(4): 593-595.
9. Lei F, Zhang H, Long Y, et al. Characteristics and phylogenetic analysis of the complete chloroplast genome of Lilium concolor Salisb.(Liliaceae) from Jilin, China[J]. Mitochondrial DNA Part B, 2022, 7(1): 30-31.
Sun C, Liu X, Zhou H, et al. The complete chloroplast genome sequence of Zanthoxylum undulatifolium Hemsl.(Rutaceae)[J]. Mitochondrial DNA Part B, 2022, 7(2): 382-384.
Sun C, Liu X, Dong P, et al. The complete chloroplast genome of Zanthoxylum piasezkii Maxim.(Rutaceae) and its phylogenetic analysis[J]. Mitochondrial DNA Part B, 2021, 6(2): 306-307.
Liu X, Sun C, Luo M, et al. The complete chloroplast genome of Zanthoxylum bungeanum var. pubescens with distinct leaf shapes[J]. Mitochondrial DNA Part B, 2021, 6(6): 1768-1769.
Li S, Wang Y P, Zhao Y F, et al. Characterization of the complete chloroplast genome of the Solanum tuberosum L. cv. Favorita (Solanaceae)[J]. Mitochondrial DNA Part B, 2021, 6(3): 909-911.
Chen S, Zhao Y, Zhang J Y, et al. Characterization of the complete chloroplast genome of the Solanum tuberosum L. cv. Shepody (Solanaceae)[J]. Mitochondrial DNA Part B, 2021, 6(8): 2342-2344.
Zhang J Y, Zhang J Y, Zhao Y F, et al. Characterization of the complete chloroplast genome of the Solanum tuberosum L. cv. Atlantic (Solanaceae)[J]. Mitochondrial DNA Part B, 2021, 6(1): 73-75.
ependent species from Peucedanum praeruptorum[J]. Physiology and Molecular Biology of Plants, 2021: 1-13.Complete chloroplast genome sequencing support Angelica decursiva is an independent species from Peucedanum praeruptorum | SpringerLink